Linear Integration of ERK Activity Predominates over Persistence Detection in Fra-1 Regulation
Taryn E. Gillies1,3
Michael Pargett1,3
Marta Minguet1
Alex E. Davies1,2
John G. Albeck1,4,*
- 1Department of Molecular and Cellular Biology, University of California, Davis, CA 95616, USA
- 2Biological Systems and Engineering Division, Lawrence Berkeley National Laboratory, Berkeley, CA 94720, USA
- 3These authors contributed equally
- 4Lead Contact
- *Correspondence: jgalbeck@ucdavis.edu
Highlights
- Multiple reporters are needed to capture the full dynamic range of ERK activity
- Fra-1 induction is related linearly to the integrated activity of ERK
- Basal transcription rate determines whether a gene may act as a persistence detector
- ERK-independent factors contribute substantially to Fra-1 variance in single cells
Graphical Abstract
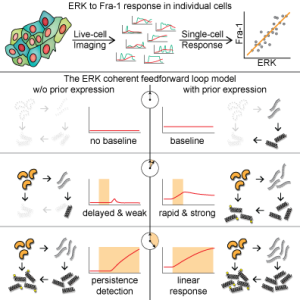
Summary
ERK signaling regulates the expression of target genes, but it is unclear how ERK activity dynamics are interpreted. Here, we investigate this question using simultaneous, live, single-cell imaging of two ERK activity reporters and expression of Fra-1, a target gene controlling epithelial cell identity. We find that Fra-1 is expressed in proportion to the amplitude and duration of ERK activity. In contrast to previous “persistence detector” and “selective filter” models in which Fra-1 expression only occurs when ERK activity persists beyond a threshold duration, our observations demonstrate that the network regulating Fra-1 expression integrates total ERK activity and responds to it linearly. However, exploration of a generalized mathematical model of the Fra-1 coherent feedforward loop demonstrates that it can perform either linear integration or persistence detection, depending on the basal mRNA production rate and protein production delays. Our data indicate that significant basal expression and short delays cause Fra-1 to respond linearly to integrated ERK activity.
Videos
ERK, Fra-1, and NCd2 response to EGF
Click here if embedded player doesn't show up
The above video shows reporter-bearing MCF-10A 5e cells responding to a 20 ng/mL EGF stimulus. Cells contain the rerporters ERKTR (red, left), Fra-1::mVenus (green, center), and NCd2 (cyan, right).
Data
The following archive contains processed cell trace data underlying this work. Data are stored as MATLAB mat files. See the accompanying PDF for further details.